01b Stitching tile folders
Use the stitching workflow when one field of view was acquired as a grid of partially overlapping image tiles. The goal is to convert many tile TIFFs into one spectral stack that can be loaded and analyzed like a normal image.
GIF placeholder: select a tile folder, parse filenames, preview the grid, and stitch.
Expected input
The stitching tab expects one folder containing tile images. Each tile filename must contain an x and y tile index that can be extracted with a regular expression.
Each tile can be:
- A 2D grayscale image.
- A 3D spectral tile in
zyxorcyxorder. - A 3D tile in
yxcorder.
For hyperspectral or multispectral tiles, choose the correct Input image order before stitching. The stitched output is returned in spectral-stack order so it can enter the normal loading and analysis workflow.
Basic workflow
- Open the HS Image Stitching tab.
- Choose the tile folder or drag a tile/folder onto the drop area.
- Set the file Pattern, for example
*.tifor*CARS*.tif. - Set the filename Regex so the table can extract x and y tile indices.
- Click Apply regex and inspect the preview table.
- Set binning, overlap, scan direction, and input order.
- Decide whether to use correlation.
- Click Stitch now.
- Inspect the stitched image.
- Save the stitched TIFF or continue directly with analysis.
The table preview is the most important diagnostic. If the grid looks wrong before stitching, the stitched image will usually also be wrong.
When to use stitching
Use the stitching workflow when:
- one field of view was acquired as many partially overlapping tiles,
- the filenames contain stable tile indices,
- the overlap is known approximately from the microscope setup,
- and you want one larger spectral stack before ROI placement or multivariate analysis.
Do not use stitching when the dataset is already one complete TIFF stack, or when the tiles are independent fields of view that should stay separate.
Pattern vs regex
The Pattern filters which files are considered:
*.tif
*THG*.tif
sample_01_*.tiff
The Regex extracts tile coordinates from the selected filenames. The regex must contain the named groups:
(?P<x>...)
(?P<y>...)
These groups tell the program which part of the filename is the x index and which part is the y index.
Default regex
The default regex is:
.*pos[_-](?P<x>-?\d+)[_-](?P<y>-?\d+).*
This matches filenames such as:
sample_pos_0_0.tif
sample_pos_1_0.tif
sample-pos-2-3.tif
Explanation:
.*ignores any text before the tile position.pos[_-]looks forpos_orpos-.(?P<x>-?\d+)captures the x index, including optional negative values.[_-]accepts_or-between x and y.(?P<y>-?\d+)captures the y index..*ignores any text after the coordinates.
Regex examples
For filenames:
tile_x03_y07.tif
use:
.*x(?P<x>\d+)_y(?P<y>\d+).*
For filenames:
scanX-1_Y-2_ch0.tif
use:
.*X(?P<x>-?\d+).*Y(?P<y>-?\d+).*
For filenames:
xyz-Table[3] - xyz-Table[7].tif
use:
.*xyz-Table\[(?P<y>\d+)\]\s*-\s*xyz-Table\[(?P<x>\d+)\].*
Here, the first table index is assigned to y and the second to x. This is allowed. The important part is that the named groups match the physical tile coordinates.
Use IGNORECASE when filename capitalization is inconsistent.
GIF placeholder: use the regex helper and show the preview table updating.
What the main settings mean
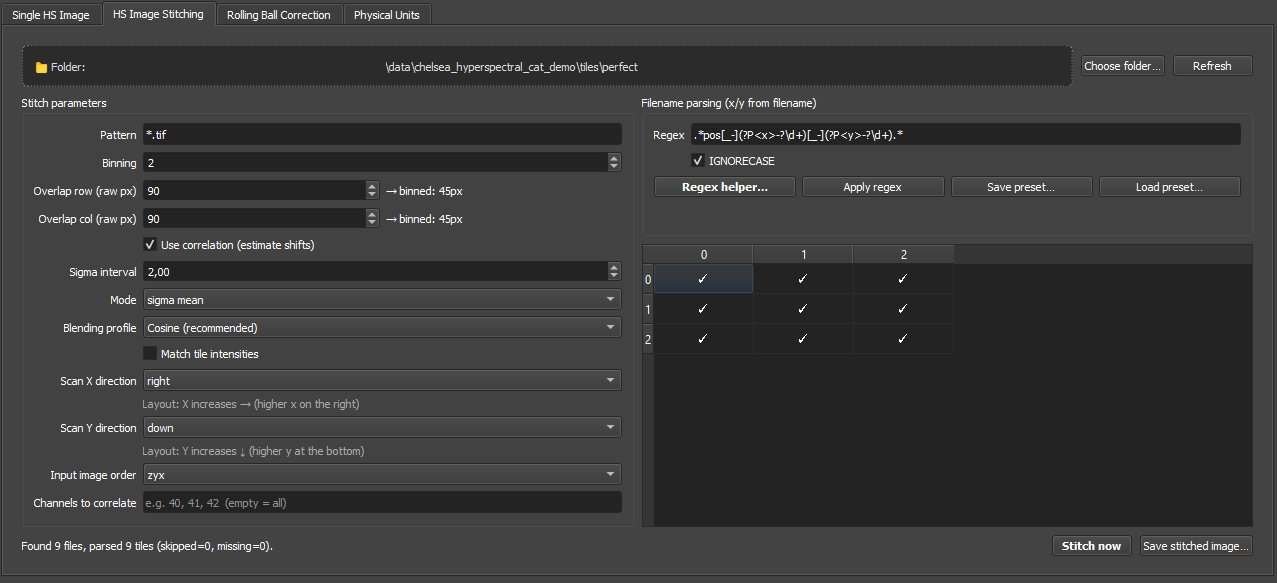
The screenshot below shows the main regions that matter for parameter choice:
- left: stitch geometry and correlation settings,
- right: filename parsing and the preview table,
- bottom: run/export actions and stitched-result status.
Pattern
Pattern is only a file filter. It decides which files in the folder are considered for parsing and stitching.
Use it when:
- one folder contains several modalities or repeated exports,
- only one subset should be stitched,
- or the folder also contains overview images, thumbnails, or metadata files.
If the preview table is empty, check Pattern before debugging the regex.
Binning and overlap
The overlap fields are entered in raw pixels:
- Overlap row (raw px): overlap between vertically adjacent tiles.
- Overlap col (raw px): overlap between horizontally adjacent tiles.
If binning is enabled, the GUI shows the effective binned overlap. Conceptually:
Example:
raw overlap = 180 px
binning = 2
effective stitched overlap = 90 px
Use the raw microscope overlap as input. Do not manually divide it before entering it into the GUI.
What these settings control:
Binning: reduces the spatial sampling before stitching. This is mainly a speed and memory setting.Overlap row (raw px): expected vertical overlap between neighboring tiles.Overlap col (raw px): expected horizontal overlap between neighboring tiles.
When to use them:
- Use
Binning = 1when precise alignment matters and the dataset is still manageable. - Increase binning when tiles are very large and the first goal is a quick preview or parameter search.
- Use the microscope’s nominal overlap as a starting point, then adjust if seams or duplicated structures remain visible.
What happens if they are wrong:
- too small overlap: duplicated structures or misregistered seams,
- too large overlap: excessive blending and spatial compression,
- too much binning: faster stitching but weaker correlation precision and less sharp seams.
How precise does the overlap value need to be?
The right strategy depends on whether you know the true overlap accurately:
- Overlap unknown or approximate (e.g. only the nominal stage value is available). Use correlation and enter an overlap that is somewhat larger than your best guess (e.g. +20–40 raw pixels). The cross-correlation search is robust enough to find the true shift within an oversized search region, and the extra margin protects you against under-shoot. Erring high in correlation mode is safer than erring low.
- Overlap known precisely (e.g. from a calibrated stage or measured directly on a reference tile). Enter the exact overlap. With correlation enabled, an oversized overlap then dilutes the score with regions that do not actually correlate, which can pull the estimated shift onto a noise peak. With correlation disabled (pure grid placement), the value sets the blend region directly, so accuracy matters even more.
Rule of thumb:
- unknown: overestimate slightly, let correlation refine.
- known: enter the exact value, do not pad.
Scan direction
Scan direction controls how the parsed x/y indices are mapped into the displayed mosaic.
Scan X direction:
right: higher x indices appear to the right.left: higher x indices appear to the left.
Scan Y direction:
down: higher y indices appear lower in the image.up: higher y indices appear higher in the image.
If the preview table or stitched result is mirrored, change the scan direction rather than changing the regex.
Use scan direction to correct orientation, not tile identification. The regex should answer "which tile is this?", while scan direction should answer "where do higher x and y indices appear in the displayed mosaic?".
Input image order
Choose the array order used inside each tile:
zyx: spectral/channel axis first, then y, then x.cyx: channel axis first, then y, then x.yxc: y, x, then channel axis.
For normal hyperspectral TIFF stacks, zyx or cyx is usually correct. For camera-style multi-channel images, yxc may be correct.
If the stitched result has swapped spatial and spectral axes, this setting is the first thing to check.
Practical rule:
zyx/cyx: choose this for spectral stacks saved frame-first.yxc: choose this for conventional image formats where channels are stored last.
If one stitched tile looks visually correct but the channel slider behaves strangely afterwards, the issue is often here.
Choosing grid placement vs correlation
There are two broad strategies:
- grid placement only: place tiles using the parsed x/y grid and the entered overlaps,
- correlation-assisted stitching: estimate small relative shifts from the image content.
Use grid placement only when:
- stage motion is reliable,
- overlap is known well,
- or the tiles have weak internal structure and correlation becomes unstable.
Use correlation when:
- the stage has small positioning errors,
- there is enough texture or contrast in the overlap region,
- and a purely geometric stitch still leaves visible seams or small jumps.
Correlation settings
When Use correlation is enabled, the program estimates small relative shifts between overlapping tiles. This is useful when the stage coordinates are approximate or the microscope has small positioning errors.
Important settings:
- Channels to correlate: leave empty to use all channels, or enter a list such as
40, 41, 42. - Mode normal: use the individual correlation offsets.
- Mode sigma: remove outlier offsets using the sigma interval.
- Mode mean: average offsets so a common offset is applied.
- Mode sigma mean: remove outliers first, then average offsets.
- Sigma interval: controls how aggressively outlier offsets are rejected.
Use a small channel list when only some channels contain reliable structure. Use all channels when the signal is broadly present and stable.
Recommended interpretation:
normal: most direct, least smoothed. Good when the correlation is already stable.mean: conservative. Good when all overlaps show about the same shift and you want one consistent correction.sigma: robust against a few bad overlaps, but still keeps local variation.sigma mean: strongest stabilization. Good default when some overlap regions are noisy or nearly empty.
How to choose Channels to correlate:
- leave it empty when most channels show similar morphology,
- restrict it when only part of the spectrum contains clear structure,
- avoid channels dominated by noise, saturation, or flat background.
How to choose Sigma interval:
- smaller values reject more offsets as outliers,
- larger values trust more measured offsets,
- if correlation seems unstable, try
sigma meanfirst before changing the overlap values.
If correlation makes the mosaic worse, disable it and use grid placement with the current blending settings.
Blending
Two controls in the stitch dialog decide how the overlapping pixels of adjacent tiles are combined into the final stitched image:
- Blending profile (dropdown):
Cosine (recommended)orLinear. - Match tile intensities (checkbox): on/off, default off.
Blending profile
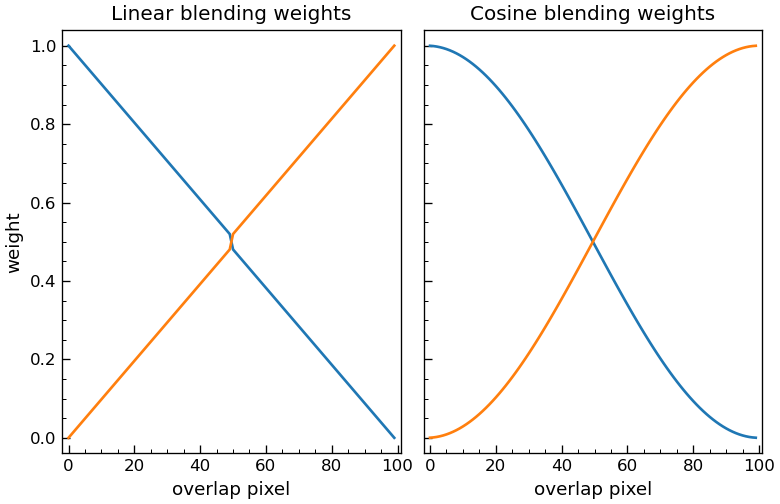
The weight ramp applied across the overlap region:
Cosine(default): a raised-cosine (Hann) curve. The two weights vary smoothly from ~1/~0 at the seam edges to 0.5/0.5 at the centre and the ramp is C¹ at the midpoint, so the human eye does not pick up the kink that a triangular ramp would leave.Linear: the legacy triangular tent. Kept available for comparison and backwards compatibility with older results.
In both cases the two weights sum to 1 at every overlap column, so the stitch is a true convex combination of the two tiles.
Match tile intensities
Each incoming tile can be scaled, per channel, so that its mean intensity inside the overlap region matches the mean of the already stitched image in the same region. This step happens before the weighted blend.
When this helps:
- visible soft brightness steps at seams caused by vignetting,
- exposure or laser-power drift between tiles,
- tile-to-tile gain differences in the detector.
When to leave it off (the default):
- quantitative work where the absolute tile intensities must be preserved exactly,
- already flat-field corrected tiles where mean intensities are guaranteed to match,
- single-tile diagnostics, where the rescaling would be a no-op anyway.
Implementation notes:
- The factor is per channel, NaN-safe, and clipped to
[0.1, 10.0]. A degenerate channel (mean ~ 0 on either side, or any non-finite intermediate) falls back to a factor of 1.0, so the step never invents or kills signal. - Only the incoming tile is rescaled; the running stitch is never modified by this step. This keeps the result reproducible regardless of the tile order, up to the natural pairwise drift of the rescaling chain.
How to choose
| Symptom | Try |
|---|---|
| Sharp kink visible at the seam centre | Cosine profile |
| Soft brightness step across the seam | Match tile intensities |
| Both | Cosine + Match tile intensities |
| Need absolute intensity preservation | Cosine + leave matching off |
Saving and reusing stitch settings
The stitching tab has its own JSON preset. This stores:
- Pattern.
- Binning.
- Raw overlap values.
- Sigma interval and mode.
- Scan x/y direction.
- Input channel order.
- Channels to correlate.
- Filename regex.
- IGNORECASE setting.
- Blending profile (cosine / linear).
- Match tile intensities (on / off).
Use stitching presets when the microscope filename pattern and tile geometry are stable across datasets.
This is especially useful when:
- the same microscope saves the same filename scheme every day,
- overlap and scan direction are fixed for one modality,
- or different users should apply the same parsing rules consistently.
Common problems
If no tiles are parsed:
- Check the file Pattern.
- Check that the regex matches the filename.
- Check that both
(?P<x>...)and(?P<y>...)exist. - Enable IGNORECASE if capitalization differs.
If the grid is mirrored:
- Change Scan X direction or Scan Y direction.
- Do not swap x and y in the regex unless the table itself shows transposed coordinates.
If seams are visible:
- Check overlap values (overestimate slightly when using correlation, set the exact value when the overlap is known).
- Check binning.
- Try correlation.
- Restrict correlation to structural channels.
- Switch the Blending profile to
Cosineif it is still onLinear. - Enable
Match tile intensitiesif the seam looks like a soft brightness step (typical for vignetting or exposure drift). Leave it off for quantitative work. - Inspect whether rolling-ball correction or normalization should be applied consistently.
If spectral channels look wrong:
- Check Input image order.
- Check that the tile TIFFs have the expected dimensions.
- Open one tile separately and confirm its channel axis.